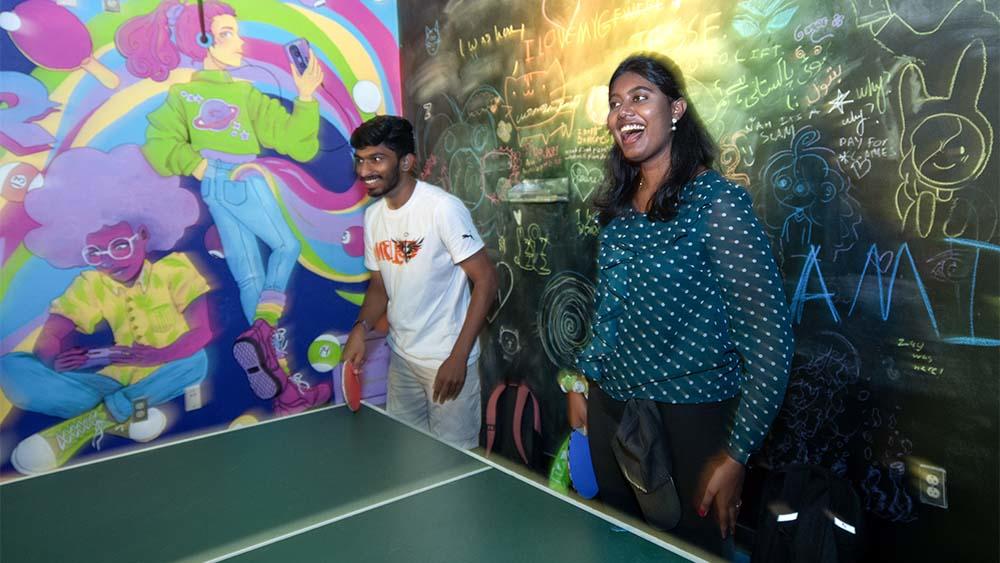
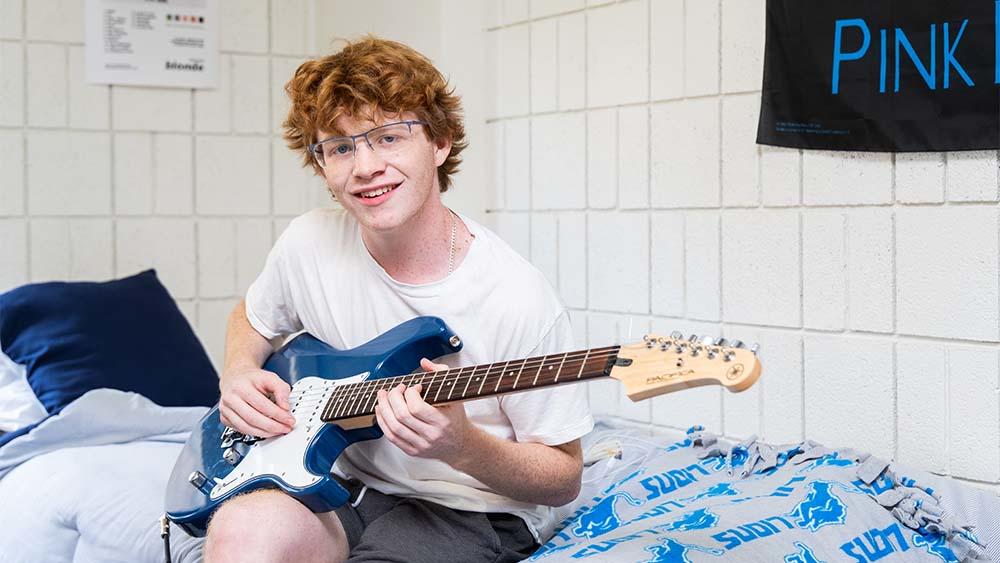
George Mason University | New Look, Same Patriot Pride | New George Mason Logo
IMPORTANT UPDATES:
-
The 2024 SPRING COMMENCEMENT will be held on May 9, 2024, at 10 am in the EagleBank Arena. Information and Tickets | Degree Celebrations Schedule | Meet Spring 2024 Commencement Speaker, Barbara Humpton
-
George Mason extends the deadline for admitted freshman student enrollment deposits to May 15. More information.
Follow Mason
Information For:
Prospective Students
Parents and Families
Current Students
Faculty and Staff
Community and Business

10 ways you can help the planet–and George Mason—this April

From first of its kind research to stories of student and alumni success, read the latest news from George Mason.

Sign up for The George, George Mason's official news source.

Events Around Campus
April 25: George Mason Wind Symphony Concert: The Blue Marble | 8 p.m. | Dewberry School of Music, Center for the Arts | Tickets
George Mason Athletics (Upcoming home events)
April 26: Baseball vs. VCU, 3 p.m.
Campus News
- April 25, 2024
- April 24, 2024
- April 22, 2024
- April 22, 2024
- April 19, 2024